Infection control
Gene accordions as potential markers for pathogenic properties
Bacteria must react to changes in the environment in order to survive. This is partly done by adapting genetic material, for example by multiplying and shortening individual genome segments. The research group led by Dr. Simon Heilbronner from the Interfaculty Institute of Microbiology and Infection Medicine at the University of Tübingen has now shown that these so-called gene accordions are frequently found in the bacterium Staphylococcus aureus and he hopes that in future they can serve as markers for the bacterium’s pathogenic properties.
Staphylococcus aureus is a widespread, spherical bacterium. It often colonises human skin and mucous membranes and is usually harmless. However, some strains are invasive, i.e. they are able to cross cell layer barriers and can cause local inflammation (abscesses) as well as life-threatening diseases such as heart inflammation or blood poisoning. These serious consequences tend to occur mostly when the host's immune system is weakened or the bacterium finds a way to escape immune defense mechanisms. The bacterium achieves the latter by adapting its genetic material. If, for example, other bacteria with advantageous genes are present in the vicinity of Staphylococcus aureus, an exchange of genetic information between the two organisms can occur (horizontal gene transfer). If this is not possible, Staphylococcus aureus bacteria can rearrange their genetic material, for example by repositioning or multiplying genome segments.
Gene accordions enable adaptation
 Electron microscope image of Staphylococcus aureus bacteria (pink). "Methicillin-Resistant Staphylococcus aureus (MRSA)" by NIAID licensed under CC BY 2.0.
Electron microscope image of Staphylococcus aureus bacteria (pink). "Methicillin-Resistant Staphylococcus aureus (MRSA)" by NIAID licensed under CC BY 2.0.Bacteria are unicellular organisms and reproduce by simple cell division, resulting in identical offspring. However, individual cells can change their genetic material by means of gene duplication and amplification (GDA). This mechanism is crucial to enable bacteria to adapt to their environment and has a major impact on their evolution. Amplification of individual genes leads to a larger amount of the gene product, i.e. the synthesised protein, which can have advantages. The number of copies in the progeny of an individual cell can vary greatly and can be up to several hundred. Depending on the environmental conditions, the gene segments expand or contract, rather like an accordion.
Dr. Simon Heilbronner's research group in Tübingen has been investigating the occurrence of gene accordions in Staphylococcus aureus for several years. When he was a doctoral student in Dublin, Heilbronner first came across this phenomenon in clinical samples from patients, which prompted him to ask the question, "Is this a mechanism that pathogens use to interact with their host?" The gene accordions detected in bacteria such as E. coli in the 1970s are mainly associated with resistance to antibiotics, heavy metal tolerance or survival in unusual conditions, but rarely with adaptation to a host. Supported by a Marie Skłodowska-Curie fellowship from the European Union, Heilbronner moved on to Tübingen to investigate where and how frequently gene accordions occur in the S. aureus genome and whether they are associated with the bacterium’s pathogenic properties.
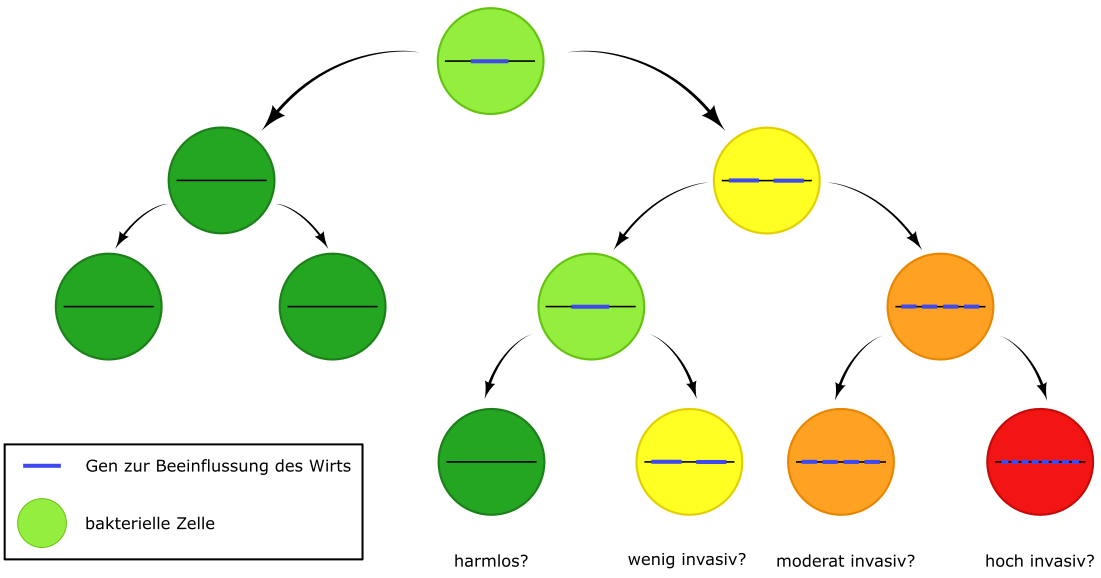 Bacteria adapt to their environment by means of gene duplication and amplification, as well as deletion, resulting in genetically different offspring. Depending on the number of copies of a gene, the offspring can have different characteristics. © Libera Lo Presti and Sophia Karner
Bacteria adapt to their environment by means of gene duplication and amplification, as well as deletion, resulting in genetically different offspring. Depending on the number of copies of a gene, the offspring can have different characteristics. © Libera Lo Presti and Sophia KarnerMany different strains of S. aureus are common in hospitals and they differ hugely in terms of pathogenicity. The basic aim of the work is to find out whether gene accordions are suitable as markers for individual pathogenic characteristics. "It would be very interesting to be able to predict how aggressive a strain is. People are fairly desperate to find such markers, which work well for predicting bacterial resistance, but too little is known about invasiveness mechanisms. We are now carrying out basic research to find out more about this," said Heilbronner, who has led his own research group at the Interfaculty Institute of Microbiology and Infection Medicine since 2018.
S. aureus has a large number of duplicated genes
 Dr. Simon Heilbronner and his research group are investigating the occurrence of gene accordions in Staphylococcus aureus. © Simon Heilbronner
Dr. Simon Heilbronner and his research group are investigating the occurrence of gene accordions in Staphylococcus aureus. © Simon HeilbronnerTo this end, Heilbronner’s team analysed the DNA sequences of almost 350 S. aureus strains that had been isolated from hospital patients. They were able to identify gene copy number variations in loci that harbour repetitive sequences, including the csa1 gene, which codes for a protein on the bacterial surface. Bacterial strains with a high csa1 copy number produce a significantly larger amount of protein. In cell culture experiments, this triggers an increased release of immunomodulatory messenger substances (for example interleukin-8), which can lead to the immune system dangerously overreacting. In the animal model, the researchers were able to confirm that the gene copy number within the csa1 gene locus changes significantly within a day after infection with S. aureus. As the bacteria divide every 30 minutes, 24 hours suffice to produce a large number of different offspring.
The exact biological function of the amplified csa1 locus is still unclear; increased invasiveness could not be proven. However, the csa1 proteins are highly immunogenic and both humans and animals produce antibodies against them. It is therefore possible that during amplification, mutations may cause increased protein variability, making it easier for the bacterium to evade the immune system.
Research on gene accordions in pathogens is still in its infancy, as they are difficult to detect using standard methods. The data published in Nature Communications in July show for the first time that this mechanism is widespread in S. aureus. Simon Heilbronner's research group is currently concentrating mainly on elucidating the biological relevance of gene accordions. "Are there regions associated with certain properties?" is one of the questions the scientists are asking. The findings may help to understand the specific adaptation of S. aureus to the host and to develop targeted drugs against it.
New cluster of excellence in Tübingen
The research is part of the Control of Microorganisms to Combat Infections cluster of excellence, which has been funded by the German federal and state governments since 2019. The cluster brings together Tübingen University and Max Planck Institute for Developmental Biology research groups from different disciplines in clinical, molecular biology and bioinformatics. The aim is to develop new forms of therapy to remove harmful bacteria from the human microbiome (i.e. the genetic material of all microbes that live in and on the human body) without having to use broad-spectrum antibiotics, as these often damage beneficial microorganisms and also promote bacterial resistance to antibiotics. "Suitable markers to determine which microorganism lines we need to get rid of altogether would be a first step towards our goal," explains Heilbronner. Gene accordions might be suitable for this purpose.
Interestingly, the researchers’ studies have shown that in S. aureus, in addition to amplification, individual gene deletions occur to an even greater extent than amplifications. This could also be a sign of environmental adaptation. In individual niches (e.g. nose, throat, armpits), certain characteristics are no longer needed and are therefore removed from the genome. In other niches, however, these particular genes may be helpful and are therefore amplified. A re-analysis of the S. aureus sequences is needed to identify such niche-specific virulence factors, taking into account the place of origin of the bacterial strains. This approach opens up further possibilities to find targets for a more precise and therefore gentler control of undesired bacteria.